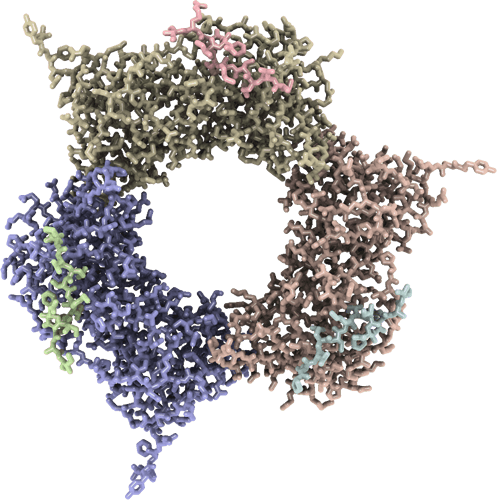
Structure of AtPCNA in complex with the PIP motif of ATXR6
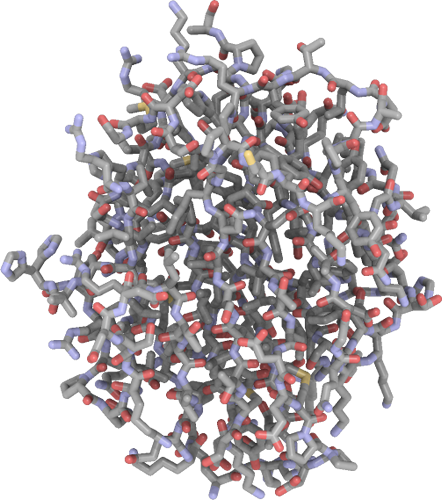
Crystal structure of Ash2L SPRY domain in complex with phosphorylated RbBP5
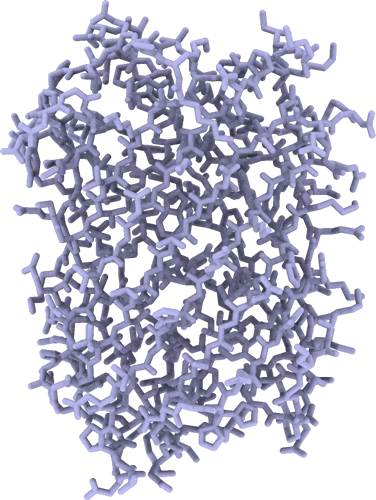
Crystal structure of mRojoA mutant – T16V – P63Y – W143G – L163V
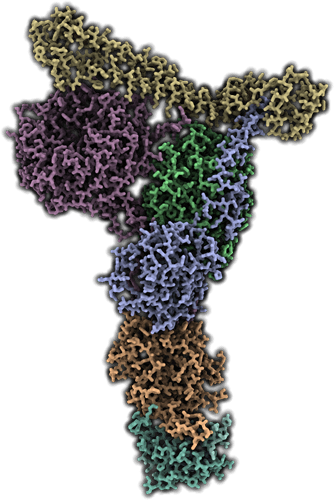
Structure of histone H3K4 methyltransferase
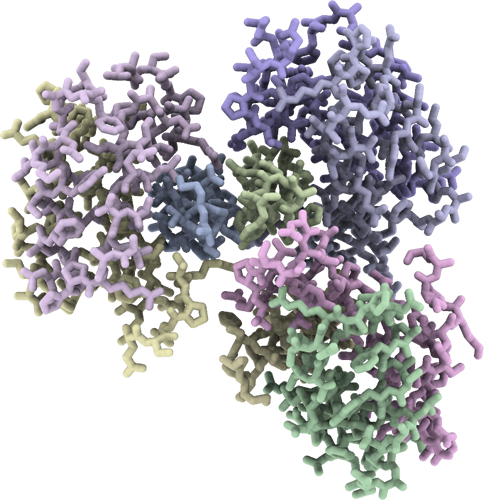
Crystal structure of DPY-30 dimerization/docking domain in complex with Ash2L Sdc1-DPY-30 Interacting region (SDI)
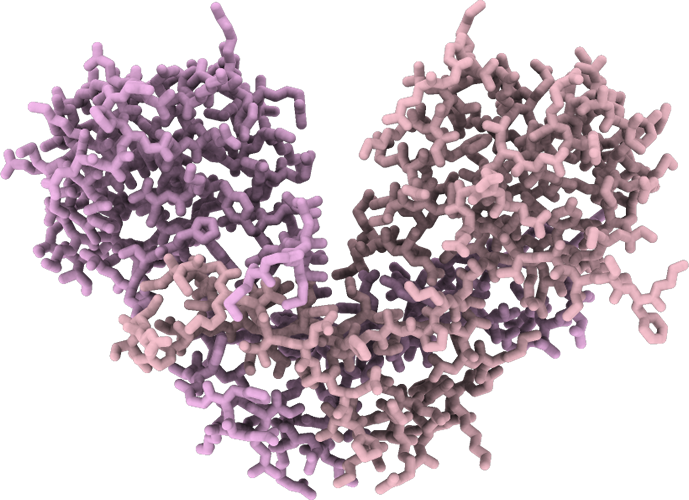
Campylobacter jejuni ferric uptake regulator S1 metalated
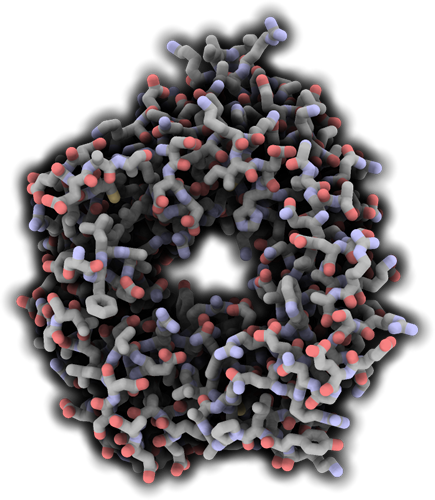
Structure of TSK/BRU1 bound to histone H3.1

PDB ID: 5VA6
Crystal structure of ATXR5 in complex with histone H3.1 mono-methylated on R26
PDB ID: 6DVS
Crystal structure of Pseudomonas stutzeri D-phenylglycine aminotransferase
Fine-tuning the stimulation of MLL1 methyltransferase activity by a histone H3 based peptide mimetic